Tyrosinase
| TYR | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | TYR, ATN, CMM8, OCA1, OCA1A, OCAIA, SHEP3, tyrosinase, Tyrosinase | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 606933; MGI: 98880; HomoloGene: 30969; GeneCards: TYR; OMA:TYR - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
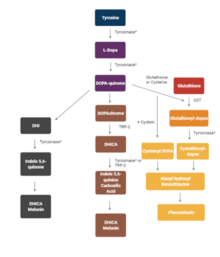
Tyrosinase is an oxidase that is the rate-limiting enzyme for controlling the production of melanin. The enzyme is mainly involved in two distinct reactions of melanin synthesis otherwise known as the Raper–Mason pathway. Firstly, the hydroxylation of a monophenol and secondly, the conversion of an o-diphenol to the corresponding o-quinone. o-Quinone undergoes several reactions to eventually form melanin.[5] Tyrosinase is a copper-containing enzyme present in plant and animal tissues that catalyzes the production of melanin and other pigments from tyrosine by oxidation. It is found inside melanosomes which are synthesized in the skin melanocytes. In humans, the tyrosinase enzyme is encoded by the TYR gene.[6]
Catalyzed reaction
[edit]Tyrosinase carries out the oxidation of phenols such as tyrosine and dopamine using dioxygen (O2). In the presence of catechol, benzoquinone is formed (see reaction below). Hydrogens removed from catechol combine with oxygen to form water.
The substrate specificity becomes dramatically restricted in mammalian tyrosinase which uses only L-form of tyrosine or DOPA as substrates, and has restricted requirement for L-DOPA as cofactor.[7]
Active site
[edit]
| monophenol monooxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Catechol-Quinone | |||||||||
| Identifiers | |||||||||
| EC no. | 1.14.18.1 | ||||||||
| CAS no. | 9002-10-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Tyrosinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Tridimensional structure of a functional unit from octopus hemocyanin | |||||||||
| Identifiers | |||||||||
| Symbol | Tyrosinase | ||||||||
| Pfam | PF00264 | ||||||||
| Pfam clan | CL0205 | ||||||||
| InterPro | IPR002227 | ||||||||
| PROSITE | PDOC00398 | ||||||||
| SCOP2 | 1hc2 / SCOPe / SUPFAM | ||||||||
| |||||||||
| Common central domain of tyrosinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Tyrosinase | ||||||||
| Pfam | PF00264 | ||||||||
| InterPro | IPR002227 | ||||||||
| PROSITE | PDOC00398 | ||||||||
| SCOP2 | 1hc2 / SCOPe / SUPFAM | ||||||||
| |||||||||
The two copper atoms within the active site of tyrosinase enzymes interact with dioxygen to form a highly reactive chemical intermediate that then oxidizes the substrate. The activity of tyrosinase is similar to catechol oxidase, a related class of copper oxidase. Tyrosinases and catechol oxidases are collectively termed polyphenol oxidases.
Structure
[edit]Tyrosinases have been isolated and studied from a wide variety of plant, animal, and fungal species. Tyrosinases from different species are diverse in terms of their structural properties, tissue distribution, and cellular location.[9] No common tyrosinase protein structure occurring across all species has been found.[10] The enzymes found in plant, animal, and fungal tissue frequently differ with respect to their primary structure, size, glycosylation pattern, and activation characteristics. However, all tyrosinases have in common a binuclear, type 3 copper centre within their active sites. Here, two copper atoms are each coordinated with three histidine residues.

Plant
[edit]In vivo, plant PPOs are expressed as about 64–68 kDa proteins consisting of three domains: a chloroplastic transit peptide (containing a ~4-9 kDa thylakoid signal peptide), a catalytically active domain (~ 37–42 kDa) containing the dinuclear copper center, and a C-terminal domain (~15–19 kDa) shielding the active site.[11]
Mammalian
[edit]Mammalian tyrosinase is a single membrane-spanning transmembrane protein.[12] In humans, tyrosinase is sorted into melanosomes[13] and the catalytically active domain of the protein resides within melanosomes. Only a small, enzymatically inessential part of the protein extends into the cytoplasm of the melanocyte.
As opposed to fungal tyrosinase, human tyrosinase is a membrane-bound glycoprotein and has 13% carbohydrate content.[14]
The derived TYR allele (rs2733832) is associated with lighter skin pigmentation in human populations. It is most common in Europe, but is also found at lower, moderate frequencies in Central Asia, the Middle East, North Africa, and among the San and Mbuti Pygmies.[15]
Bacterial
[edit]In peatlands, bacterial tyrosinases are proposed to act as key regulators of carbon storage by removing phenolic compounds, which inhibit the degradation of organic carbon.[16]
Fungal
[edit]In the fungus Neurospora crassa, four different forms of tyrosinase were distinguished among different strains.[17] In each strain only one structure-determining genetic region was found for the enzyme.
Gene regulation
[edit]The gene for tyrosinase is regulated by the microphthalmia-associated transcription factor (MITF).[18][19]

Clinical significance
[edit]A mutation in the tyrosinase gene resulting in impaired tyrosinase production leads to type I oculocutaneous albinism, a hereditary disorder that affects one in every 20,000 people.[21]
Tyrosinase activity is very important. If uncontrolled during the synthesis of melanin, it results in increased melanin synthesis. Decreasing tyrosinase activity has been targeted for the improvement or prevention of conditions related to the hyperpigmentation of the skin, such as melasma and age spots.[22]
Several polyphenols, including flavonoids or stilbenoid, substrate analogues, free radical scavengers, and copper chelators, have been known to inhibit tyrosinase.[23] Henceforth, the medical and cosmetic industries are focusing research on tyrosinase inhibitors to treat skin disorders.[5]
Inhibitors
[edit]Known Tyrosinase inhibitors are the following:[24]
Genetics
[edit]While albinism is common, there have only been a few studies about the genetic mutations in the tyrosinase genes of animals. One of them was on Bubalus bubalis (water buffalo). The tyrosinase mRNA sequence of the wild-type B. bubalis is 1,958 base pairs (bp) with an open reading frame (ORF) of 1,593 bp long, which translates to 530 amino acids. Meanwhile, the tyrosinase gene of the albino B. bubalis (GenBank JN_887463) is truncated at position 477, caused by a point mutation in nucleotide 1431 which converts a Tryptophan (TGG) into a stop codon (TGA), resulting in a shorter and inactive tyrosinase gene.[25] Other albinos have point mutations that appear to inactivate Tyrosinase without truncation (see table and figure for examples).
| Species | Common name | Amino Acid mutation | GenBank | Uniprot ID |
|---|---|---|---|---|
| Bubalus bubalis | Water Buffalo | W477 -> Stop codon | JN_887462 | J7FBF2 |
| Pelophylax nigromaculatus | Pond Frog | Deletion of a K228 | Q04604 | |
| Glandirana rugosa | Wrinkled Frog | G376 -> D376 | A0A1I9FZH0 | |
| Fejervarya kawamurai | Rice Frog | G57 -> R57 | A0A1E1G7U0 |

Knowing that there are a few studies about the genomic data of the tyrosinase gene, there are only a handful of studies on the mutations in albino amphibians. Miura et al. (2018) investigates the amino acid mutations in the tyrosinase gene in three albino frogs: Pelophylax nigromaculatus (pond frog), Glandirana rugosa (wrinkled frog) and Fejervarya kawamurai (rice frog). In total, five different populations were studied of which three were P. nigromaculatus and one each of G. rugosa and F. kawamurai. In two of the three P. nigromaculatus populations, there was a frameshift mutation because of the insertion of a thymine within exons 1 and 3, and the third population lacked three nucleotides that encoded a Lysine in exon 1. The population of G. rugosa had a missense mutation where there was an amino acid substitution from a Glycine to Aspartic acid, and the mutation of F. kawamurai was also an amino acid substitution from Glycine to Arginine. The mutation for G. rugosa and F. kawamurai occurs in exons 1 and 3. The mutations of the third population of P. nigromaculatus, and the mutations of G. rugosa and F. kawamurai occurred in areas that are highly conserved among vertebrates which could result in a dysfunctional tyrosinase gene.[26]


Evolution
[edit]
Tyrosinase is a highly conserved protein in animals and apparently arose already in bacteria. The tyrosinase related protein (Tyrp1) and dopachrome tautomerase (Dtc), which encode for protein implicated in melanin synthesis which are the common regulatory elements of exon/intron structure. The development of the three types of vertebrate pigment cells, although different, thus converge at a certain point to allow the expression of members of the tyrosinase family, in order to produce melanin pigments.[28] Tyrosinase family related genes plays an important role in the evolution, genetics, and developmental biology of pigment cells, as well as to approach human disorders associated with defects in their synthesis, regulation or function in vertebrates three types of melanin producing pigment cells are well known since embryonic origin i.e., from the neural crest, neural tube and pineal body. All of them have the capacity to produce melanin pigments. Their biosynthesis is governed by evolutionary conserved enzymes of the tyrosinase family( tyr, tyr1 and tyr2) also called DOPAchrome tautomerase (dct). Among them Tyr plays significance role in melanin production. However, sequenced genome from the different taxa for evolutionary analysis in the depth become more crucial in present study.[29] Similarly, the type-3 copper protein family perform various biological function including pigment formation, innate immunity and oxygen transport. The combine genetic phylogenetic and structural analysis concluded that the original type-3 copper protein possessed a single peptide and grouped into α subclass. The ancestral protein gene underwent to two duplication i.e., first one prior to divergence of unknown eukaryotic lineage and second one before diversification. The prior duplication gave rise to cytosolic form(β) and latter duplication gave membrane bound form (Γ). The structural comparison concluded that active site of α and γ forms are covered by aliphatic amino acids and β form covered with aromatic residue. Thus, the evolution of these gene family is the lineage of multicellular eukaryotes due to loss of one or more of these three subclasses and lineage-specific expansion of one or both of the remaining subclasses.[30] The genomic conserved nucleotide alignments of the tyrosinase among the vertebrate family like frogs, snakes and human suggests that it has evolved from one ancestral tyrosinase gene. The duplication and mutation of this gene is probably responsible for the emergence of a tyrosinase-related gene.[31]
Applications
[edit]In the food industry
[edit]In the food industry, tyrosinase inhibition is desired as tyrosinase catalyzes the oxidation of phenolic compounds found in fruits and vegetables into quinones, which gives an undesirable taste and color and also decreases the availability of certain essential amino acids as well as the digestibility of the products. As such, highly effective tyrosinase inhibitors are also needed in agriculture and the food industry.[14] Well known tyrosinase inhibitors include kojic acid,[32] tropolone,[33] coumarins,[34] vanillic acid, vanillin, and vanillic alcohol.[35]
In the cosmetic industry
[edit]Lighter skin complexion has been associated with youth and beauty across various Asian cultures. Recent research by cosmetic companies has been focused on the development of novel whitening agents that selectively suppress tyrosinase activity to reduce hyperpigmentation while avoiding cytotoxicity of healthy melanocytes.[36] Traditional pharmacological agents such as corticosteroids, hydroquinone, and amino numeric chloride lighten skin through the inhibition of melanocyte maturation.[37] However, these agents are associated with adverse effects. Cosmetic companies have been focused on developing novel whitening agents that selectively suppress the activity of tyrosinase to reduce hyperpigmentation while avoiding melanocyte cytotoxicity as tyrosinase is the rate-limiting step of the melanogenesis pathway.
In insects
[edit]Tyrosinase has a wide range of functions in insects, including wound healing, sclerotization, melanin synthesis and parasite encapsulation. As a result, it is an important enzyme as it is the defensive mechanism of insects. Some insecticides are aimed to inhibit tyrosinase.[14]
In mussel-glue inspired polymers
[edit]Tyrosinase activated polymerization of peptides, containing cysteine and tyrosine residues, lead to mussel-glue inspired polymers. The tyrosine residues are enzymatically oxidized to dopaquinones, to which thiols of cysteine could link by an intermolecular Michael-addition. The resulting polymers adsorb strongly to various surfaces with high adhesion energies.[38][39]
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000077498 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000004651 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Kumar CM, Sathisha UV, Dharmesh S, Rao AG, Singh SA (Mar 2011). "Interaction of sesamol (3,4-methylenedioxyphenol) with tyrosinase and its effect on melanin synthesis". Biochimie. 93 (3): 562–9. doi:10.1016/j.biochi.2010.11.014. PMID 21144881.
- ^ Barton DE, Kwon BS, Francke U (Jul 1988). "Human tyrosinase gene, mapped to chromosome 11 (q14----q21), defines second region of homology with mouse chromosome 7". Genomics. 3 (1): 17–24. doi:10.1016/0888-7543(88)90153-X. PMID 3146546.
- ^ Hearing VJ, Ekel TM, Montague PM, Nicholson JM (Feb 1980). "Mammalin tyrosinase. Stoichiometry and measurement of reaction products". Biochimica et Biophysica Acta (BBA) - Enzymology. 611 (2): 251–68. doi:10.1016/0005-2744(80)90061-3. PMID 6766744.
- ^ PDB: 1WX3; Matoba Y, Kumagai T, Yamamoto A, Yoshitsu H, Sugiyama M (2006). "Crystallographic evidence that the dinuclear copper center of tyrosinase is flexible during catalysis". J. Biol. Chem. 281 (13): 8981–8990. doi:10.1074/jbc.M509785200. PMID 16436386.
- ^ Mayer AM (Nov 2006). "Polyphenol oxidases in plants and fungi: going places? A review". Phytochemistry. 67 (21): 2318–31. Bibcode:2006PChem..67.2318M. doi:10.1016/j.phytochem.2006.08.006. PMID 16973188.
- ^ Jaenicke E, Decker H (Apr 2003). "Tyrosinases from crustaceans form hexamers". The Biochemical Journal. 371 (Pt 2): 515–23. doi:10.1042/BJ20021058. PMC 1223273. PMID 12466021.
- ^ Mayer AM (November 2006). "Polyphenol oxidases in plants and fungi: going places? A review". Phytochemistry. 67 (21): 2318–31. Bibcode:2006PChem..67.2318M. doi:10.1016/j.phytochem.2006.08.006. PMID 16973188.
- ^ Kwon BS, Haq AK, Pomerantz SH, Halaban R (Nov 1987). "Isolation and sequence of a cDNA clone for human tyrosinase that maps at the mouse c-albino locus". Proceedings of the National Academy of Sciences of the United States of America. 84 (21): 7473–7. Bibcode:1987PNAS...84.7473K. doi:10.1073/pnas.84.21.7473. PMC 299318. PMID 2823263.
- ^ Theos AC, Tenza D, Martina JA, Hurbain I, Peden AA, Sviderskaya EV, Stewart A, Robinson MS, Bennett DC, Cutler DF, Bonifacino JS, Marks MS, Raposo G (Nov 2005). "Functions of adaptor protein (AP)-3 and AP-1 in tyrosinase sorting from endosomes to melanosomes". Molecular Biology of the Cell. 16 (11): 5356–72. doi:10.1091/mbc.E05-07-0626. PMC 1266432. PMID 16162817.
- ^ a b c Kim YJ, Uyama H (Aug 2005). "Tyrosinase inhibitors from natural and synthetic sources: structure, inhibition mechanism and perspective for the future". Cellular and Molecular Life Sciences. 62 (15): 1707–23. doi:10.1007/s00018-005-5054-y. PMC 11139184. PMID 15968468. S2CID 8280251.
- ^ "Allele Frequency For Polymorphic Site: rs2733832". ALFRED. Archived from the original on 20 August 2016. Retrieved 23 June 2016.
- ^ Panis F, Krachler RF, Krachler R, Rompel A (June 2021). "Expression, Purification, and Characterization of a Well-Adapted Tyrosinase from Peatlands Identified by Partial Community Analysis". Environmental Science & Technology. 55 (16): 11445–11454. Bibcode:2021EnST...5511445P. doi:10.1021/acs.est.1c02514. PMC 8375020. PMID 34156250.
- ^ HOROWITZ NH, FLING M, MACLEOD H, SUEOKA N (August 1961). "A genetic study of two new structural forms of tyrosinase in Neurospora". Genetics. 46 (8): 1015–24. doi:10.1093/genetics/46.8.1015. PMC 1210246. PMID 13715943.
- ^ Hou L, Panthier JJ, Arnheiter H (Dec 2000). "Signaling and transcriptional regulation in the neural crest-derived melanocyte lineage: interactions between KIT and MITF". Development. 127 (24): 5379–89. doi:10.1242/dev.127.24.5379. PMID 11076759.
- ^ Hoek KS, Schlegel NC, Eichhoff OM, Widmer DS, Praetorius C, Einarsson SO, Valgeirsdottir S, Bergsteinsdottir K, Schepsky A, Dummer R, Steingrimsson E (Dec 2008). "Novel MITF targets identified using a two-step DNA microarray strategy". Pigment Cell & Melanoma Research. 21 (6): 665–76. doi:10.1111/j.1755-148X.2008.00505.x. PMID 19067971.
- ^ M. S. Eller, K. Ostrom, and B. A. Gilchrest, “DNA damage enhances melanogenesis,” Proceedings of the National Academy of Sciences of the United States of America, vol. 93, no. 3, pp. 1087–1092, 1996
- ^ Witkop CJ (Oct 1979). "Albinism: hematologic-storage disease, susceptibility to skin cancer, and optic neuronal defects shared in all types of oculocutaneous and ocular albinism". The Alabama Journal of Medical Sciences. 16 (4): 327–30. PMID 546241.
- ^ Ando H, Kondoh H, Ichihashi M, Hearing VJ (Apr 2007). "Approaches to identify inhibitors of melanin biosynthesis via the quality control of tyrosinase". The Journal of Investigative Dermatology. 127 (4): 751–61. doi:10.1038/sj.jid.5700683. PMID 17218941.
- ^ Chang TS (Jun 2009). "An updated review of tyrosinase inhibitors". International Journal of Molecular Sciences. 10 (6): 2440–75. doi:10.3390/ijms10062440. PMC 2705500. PMID 19582213.
- ^ Pillaiyar T, Manickam M, Namasivayam V (December 2017). "Skin whitening agents: medicinal chemistry perspective of tyrosinase inhibitors". Journal of Enzyme Inhibition and Medicinal Chemistry. 32 (1): 403–425. doi:10.1080/14756366.2016.1256882. PMC 6010116. PMID 28097901.
- ^ Damé, Maria Cecília Florisbal; Xavier, Gildenor Medeiros; Oliveira-Filho, José Paes; Borges, Alexandre Secorun; Oliveira, Henrique Nunes; Riet-Correa, Franklin; Schild, Ana Lucia (2012-07-20). "A nonsense mutation in the tyrosinase gene causes albinism in water buffalo". BMC Genetics. 13: 62. doi:10.1186/1471-2156-13-62. ISSN 1471-2156. PMC 3411452. PMID 22817390.
- ^ a b Miura, Ikuo; Tagami, Masataka; Fujitani, Takeshi; Ogata, Mitsuaki (2018-02-10). "Spontaneous tyrosinase mutations identified in albinos of three wild frog species". Genes & Genetic Systems. 92 (4): 189–196. doi:10.1266/ggs.16-00061. ISSN 1880-5779. PMID 28674275.
- ^ Kim, Young-Hyun; Park, Sang-Je; Choe, Se-Hee; Lee, Ja-Rang; Cho, Hyeon-Mu; Kim, Sun-Uk; Kim, Ji-Su; Sim, Bo-Woong; Song, Bong-Seok; Lee, Youngjeon; Jin, Yeung Bae; Hong, Jung-Joo; Jeong, Kang-Jin; Kang, Philyong; Baek, Seung-Ho; Lee, Sang-Rae; Huh, Jae-Won; Chang, Kyu-Tae (2017). "Identification and characterization of the tyrosinase gene ( TYR ) and its transcript variants ( TYR_1 and TYR_2 ) in the crab-eating macaque ( Macaca fascicularis )". Gene. 630: 21–27. doi:10.1016/j.gene.2017.07.047. PMID 28756020.
- ^ Agnes Camacho-Hubner, Christelle Richard, Friedrich Beermann (20 Feb 2022). "Genomic structure and evolutionary conservation of the tyrosinase gene family from Fugu". Gene. 285 (1–2): 59–68. doi:10.1016/s0378-1119(02)00411-0. PMID 12039032.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Rosaria Esposito, Salvatore D'Aniello, Paola Squarzoni, Maria Rosa Pezzotti, Filomena Ristoratore and Antonietta Spagnuolo (2012). "New Insights into the Evolution of Metazoan Tyrosinase Gene Family". PLOS ONE. 7 (4): e35731. Bibcode:2012PLoSO...735731E. doi:10.1371/journal.pone.0035731. PMC 3334994. PMID 22536431.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Aguilera, Felipe; McDougall, Carmel; Degnan, Bernard M. (2013-05-01). "Origin, evolution and classification of type-3 copper proteins: lineage-specific gene expansions and losses across the Metazoa". BMC Evolutionary Biology. 13 (1): 96. Bibcode:2013BMCEE..13...96A. doi:10.1186/1471-2148-13-96. ISSN 1471-2148. PMC 3658974. PMID 23634722.
- ^ F. Murisier and F. Beermann (2006). "Genetics of pigment cells:lessons from the tyrosinase gene family". Histology and Histopathology. 21 (5): 567–578. PMID 16493586.
- ^ Mendes E, Perry Mde J, Francisco AP (May 2014). "Design and discovery of mushroom tyrosinase inhibitors and their therapeutic applications". Expert Opinion on Drug Discovery. 9 (5): 533–54. doi:10.1517/17460441.2014.907789. PMID 24708040. S2CID 12589166.
- ^ Rescigno A, Sollai F, Pisu B, Rinaldi A, Sanjust E (Aug 2002). "Tyrosinase inhibition: general and applied aspects". Journal of Enzyme Inhibition and Medicinal Chemistry. 17 (4): 207–18. doi:10.1080/14756360210000010923. PMID 12530473.
- ^ Sollai F, Zucca P, Sanjust E, Steri D, Rescigno A (December 2008). "Umbelliferone and esculetin: inhibitors or substrates for polyphenol oxidases?". Biological & Pharmaceutical Bulletin. 31 (12): 2187–93. doi:10.1248/bpb.31.2187. hdl:11584/105440. PMID 19043197.
- ^ Rescigno A, Casañola-Martin GM, Sanjust E, Zucca P, Marrero-Ponce Y (March 2011). "Vanilloid derivatives as tyrosinase inhibitors driven by virtual screening-based QSAR models". Drug Testing and Analysis. 3 (3): 176–81. doi:10.1002/dta.187. PMID 21125547.
- ^ Qian, W., Liu, W., Zhu, D., Cao, Y., Tang, A., Gong, G., Su, H."Natural skin‑whitening compounds for the treatment of melanogenesis (Review)". Experimental and Therapeutic Medicine 20.1 (2020): 173-185.
- ^ Lajis AFB and Ariff AB: Discovery of new depigmenting compounds and their efficacy to treat hyperpigmentation: Evidence from in vitro study. J Cosmet Dermatol. 18:703–727. 2019.
- ^ Horsch J, Wilke P, Pretzler M, Seuss M, Melnyk I, Remmler D, et al. (November 2018). "Polymerizing Like Mussels Do: Toward Synthetic Mussel Foot Proteins and Resistant Glues". Angewandte Chemie. 57 (48): 15728–15732. doi:10.1002/anie.201809587. PMC 6282983. PMID 30246912.
- ^ Arias S, Amini S, Horsch J, Pretzler M, Rompel A, Melnyk I, et al. (October 2020). "Toward Artificial Mussel-Glue Proteins: Differentiating Sequence Modules for Adhesion and Switchable Cohesion". Angewandte Chemie. 59 (42): 18495–18499. doi:10.1002/anie.202008515. PMC 7590116. PMID 32596967.
External links
[edit]- GeneReviews/NCBI/NIH/UW entry on Oculocutaneous Albinism Type 1
- Tyrosinase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)





